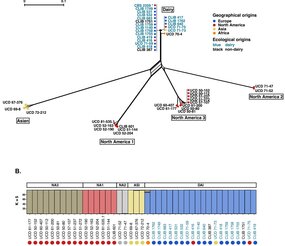
Contrasting genomic evolution between domesticated and wild Kluyveromyces lactis yeast populations.
Anne Friedrich, Jean-Sébastien Gounot, Andreas Tsouris, Claudine Bleykasten, Kelle Freel, Claudia Caradec, Joseph Schacherer.
Genome Biology and Evolution. 2023. doi:10.1093/gbe/evad004
The process of domestication has variable consequences on genome evolution leading to different phenotypic signatures. Access to the complete genome sequences of a large number of individuals makes it possible to explore the different facets of this domestication process. Here, we sought to explore the genome evolution of Kluyveromyces lactis, a yeast species well-known for its involvement in dairy processes but also present in natural environments. Using a combination of short and long-read sequencing strategies, we investigated the genomic variability of 41 K. lactis isolates and found that the overall genetic diversity of this species is very high (θw = 3.3 × 10−2) compared to other species such as Saccharomyces cerevisiae (θw = 1.6 × 10−2). However, the domesticated dairy population shows a reduced level of diversity (θw = 1 × 10−3), probably due to a domestication bottleneck. In addition, this entire population is characterized by the introgression of the LAC4 and LAC12 genes, responsible for lactose fermentation and coming from the closely related species, Kluyveromyces marxianus, as previously described. Our results also highlighted that the LAC4/LAC12 gene cluster was acquired through multiple and independent introgression events. Finally, we also identified several genes that could play a role in adaptation to dairy environments through copy number variation. These genes are involved in sugar consumption, flocculation and drug resistance, and may play a role in dairy processes. Overall, our study illustrates contrasting genomic evolution and sheds new light on the impact of domestication processes on it.